
|
Panagene supplies flexible designs of PNA oligomers for synthesis by taking advantages of its proprietary technology. All PNA oligomers are manufactured according to ISO9001/2000 and provided with report of HPLC and MALDI-TOF analysis.
When you decide base sequence, linker, label and modification of PNA oligomers, submit your order by online at Order & Inquiry. You can also submit your order through email or Fax. When any technical support in design of your PNA oligomers is needed, we provide our knowledge and experiences related with PNA synthesis and application.
|

|
|
Panagene supplies PNA oligomers with any further modifications at N-terminus at our customer's request.
|

|
|
N-terminal is capped by acetyl group to block any further reactivity.
|

|
|
To increase the solubility of PNA and to avoid self-aggregation, either or both termini may be modified with lysine, O-linker or other solubility enhancing molecules. To apply on PNA-based microarray or other format, N-terminal group can be modified to appropriate functional groups such as carboxylic acid and thiol. Panagene also supplies any further PNA modifications that are not listed here.
|

|
|
Use of appropriate spacers such as O-linker is recommended to avoid an adverse effect by a reporter in hybridization process. Although direct labeling at N-terminus is usually possible.
|

|
|
Additional use of lysine or cysteine residue is recommended since labeling is carried out via lysine /cysteine residue at C-terminus.
|

|
|
Panagene also supplies custom-based dual- or mutli-labeled PNA oligomers that can be used for molecular beacons and other uses as per request.
|

|
|
Label
|
Ąëabs(nm)
|
Ąëem(nm)
|
Label
|
Ąëabs(nm)
|
Ąëem(nm)
|
|
FAM, FITC
|
492
|
518
|
BHQ-1
|
480-580
|
-
|
|
TAMRA
|
553
|
576
|
BHQ-2
|
560-670
|
-
|
|
TexasRed
|
592
|
614
|
Dabcyl
|
454
|
-
|
|
Thiazol Orange
|
510
|
530
|
Digoxin
|
-
|
-
|
|
Cy3
|
550
|
570
|
Biotin
|
-
|
-
|
|
Cy5
|
650
|
670
|
|
|
|
|
JOE
|
520
|
548
|
|
|
|
|
ATTO425
|
436
|
484
|
|
|
|
|
ATTO550
|
554
|
576
|
|
|
|
|
|
Other fluorescence labelings are also served as per request. Please contact us for technical discussion .
|

|
|
No
|
Linker
|
Structure
|
|
1
|
O
|

|
|
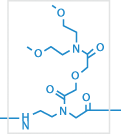
2
|
E
|
|
|
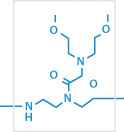
3
|
X
|
|
|

|
PNA molecules can be chemically linked to peptide to improve physical properties such as cellular uptake and solubility. There are many examples of CPPs (Cell Penetrating Peptides)-conjugated PNAs used in a variety of applications.
Generally, PNA-peptide conjugates are sequentially synthesized on resin from C-terminus to N-terminus.But in some cases, PNA and peptide can be linked with special linkage such as disulfide or SMCC/thiol.
Panagene also supplies PNA oligomers conjugated not only with well-known CPPs but with your proprietary peptide if needed.
|

|
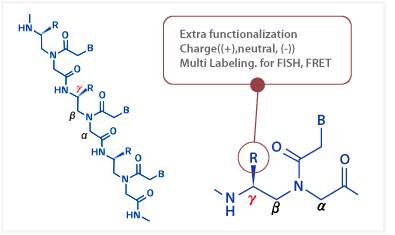
|
ĄăPNA has a stereogenic center at the Ąă-carbon atom of the N-aminoethylglycine unit. Compared to conventional PNAs, the Ąă-substituted PNAs have several advantages such as improved solubility, less self-aggregation, increased stability of PNA-DNA duplexes, and flexibility for multi labeling and other functionalization. Therefore, ĄăPNA has great potential in its applications for both diagnostics and therapeutics.
Panagene supplies oligomers containing ĄăPNAs for your research and development purposes.
** Available moiety at gamma
ˇ¤ Lysine
ˇ¤ Alanine
ˇ¤ Glutamic acid
ˇ¤ miniPEG
** Contact us for other moieties.
|

|
 Opportunities for extra functionalization at the Ąă position Opportunities for extra functionalization at the Ąă position
|
 Labeling flexibility (dual/multi labeling to backbone) Labeling flexibility (dual/multi labeling to backbone)
|
 Extra handle for improved cell penetration Extra handle for improved cell penetration
|
 Extra handle for improved water solubility Extra handle for improved water solubility
|
 Less self-aggregation due to the pre-organized Ąá-helical structure Less self-aggregation due to the pre-organized Ąá-helical structure
|
 Application for FISH probe due to less self-aggregation Application for FISH probe due to less self-aggregation
|
 Higher binding affinity to the complementary DNA or RNA than conventional PNAs Higher binding affinity to the complementary DNA or RNA than conventional PNAs
|

|
|
Criteria
|
ĄăPNA
|
PNA
|
|
Backbone modification
|
possible
|
impossible
|
|
Labeling flexibility on backbone
|
O
|
X
|
|
Water solubility
|
Extra handle for improved water solubility
|
No extra handle
|
|
Hybridization affinity with DNA
|
At least 2 ˇÉ higher Tm per modified base
|
-
|
|
Tm for each single mismatch
|
Lowering 18ˇÉ
|
Lowering 15ˇÉ
|
|
Chemical stability
|
Stable
|
Stable
|
|
Thermal stability
|
Good
|
Good
|
|
|
|

|
|
|

|
 PNA oligomers can form duplexes in either orientation, but the anti-parallel orientation is strongly preferred for all antisense and PNA oligomers can form duplexes in either orientation, but the anti-parallel orientation is strongly preferred for all antisense and
DNA probe type applications. The N-terminal of the PNA oligomer is equivalent to the 5ˇŻ-end of an oligonucleotide and is often
referred to as ˇ®5ˇŻ-end of PNAˇ±.
|
|
 PNA/DNA-duplex generally has higher Tm than corresponding DNA/DNA-duplex. Roughly, there is increase of about 1ˇĆC per PNA/DNA-duplex generally has higher Tm than corresponding DNA/DNA-duplex. Roughly, there is increase of about 1ˇĆC per
base pair at 100 mM NaCl in Tm value.
|
|
|
Sequence
|
Tm value (°C)
|
|
DNA/DNA
|
DNA/PNA
|
|
TAA CGG TAT TTG GAG(15-mer)
|
56
|
71
|
|
|
 Due to high binding affinity to its complementary DNA sequences, designs of long PNA oligomers are usually not needed for Due to high binding affinity to its complementary DNA sequences, designs of long PNA oligomers are usually not needed for
hybridization. In general, PNA oligomers with 12-18 bases are recommended. However, PNA oligomers with 27 bases can be
also supplied as per request.
|
|
 It is strongly recommended to avoid any self-complementary sequences such as inverse repeats, hairpin forming and It is strongly recommended to avoid any self-complementary sequences such as inverse repeats, hairpin forming and
palindromic sequences because PNA/PNA interaction is stronger than PNA/DNA interaction.
|
|
 It is also recommended to avoid purine-rich sequences because purine-rich PNA oligomers tend to be aggregated and then It is also recommended to avoid purine-rich sequences because purine-rich PNA oligomers tend to be aggregated and then
result in low solubility in aqueous solution. Please limit the purine contents up to 80%. Lower than 60% is more desirable.
|
In general, the maximum purine stretch should not exceed five bases in a row. However, depending on PNA
sequence, longer purine stretch longer than fives bases may be acceptable.
|
|
 For enhancing solubility of PNA probes, addition of solubility enhancer is recommended. For enhancing solubility of PNA probes, addition of solubility enhancer is recommended.
e.g.: O linker (eg1 linker), E linker, X linker, or Lysine
|
|
 For labeling at N-terminus, insertion of two O linkers between PNA and fluorescent label is recommended. For labeling at N-terminus, insertion of two O linkers between PNA and fluorescent label is recommended.
|
|
 For labeling at C-terminus, insertion of one lysine or cysteine between PNA and fluorescent label is needed. For labeling at C-terminus, insertion of one lysine or cysteine between PNA and fluorescent label is needed.
|
|