|
PANAMutyper™ R EGFR
|
|
Lung cancer is a metastatic cancer which the cancer cells are developed in the bronchial tubes or alveoli and grow along the blood vessel or lymphatic vessel. There are two types of lung cancer based on the sizes and shape of the cancer cell; non-small cell lung cancer (NSCLC) takes 75% and small cell lung cancer takes 25%. Usually, it is hard to find lung cancer in the early stage and is diagnosed as lung cancer when it is already progressed. So the prognosis tends to be bad. It is known that the patients with EGFR mutation show very high drug responses against EGFR tyrosine kinase inhibitor (TKI) such as Gefitinib (Iressa, AstraZenca) and Erlotinib (Tarceba, Roche). We expect that genotyping EGFR genes of the lung cancer patients enables predicting drug response before treatment and this will lead effective lung cancer treatment.
|
|
PANAMutyper™ R EGFR |
| Product Name |
|
Cat No. |
|
Size
|
|
PANAMutyper™ R EGFR (47 Mutations) |
|
PNAR-3001 |
|
24 tests |
|
|
|
|
|
|
|
EGFR Mutation Details |
| No. |
Reagent |
Exon |
Amino Acid Change |
Nucleotide change |
Cosmic No. |
| 1 |
G719 |
E18 |
G719A |
c.2156G>C |
6239 |
| 2 |
G719S |
c.2155G>A |
6252 |
| 3 |
G719C |
c.2155G>T |
6253 |
| 4 |
E19del A |
19 |
p.E746_A750del |
c.2235_2249 del 15 |
6223 |
| 5 |
p.E746_A750del |
c.2236_2250 del 15 |
6225 |
| 6 |
p.E746_T751>IP |
c.2235_2251>AATTC |
13552 |
| 7 |
p.K745_E749del |
c.2233_2247 del 15 |
26038 |
| 8 |
p.E746_T751>I |
c.2235_2252 AAT (complex) |
13551 |
| 9 |
p.E746_A750del |
c.2235_2248>AATTC |
13550 |
| 10 |
p.L747_S752>Q |
c.2239_2256>CAA |
12403 |
| 11 |
p.S752_I759del |
c.2253_2276 del 24 |
12416 |
| 12 |
p.E746_T751>VA |
c.2237_2253>TTGCT |
12422 |
| 13 |
p.E746_T751>A |
c.2237_2251 del 15 |
12678 |
| 14 |
p.L747_T751del |
c.2239_2253 del 15 |
6254 |
| 15 |
p. L747_T751del |
c.2238_2252 del 15 |
23571 |
| 16 |
p.L747_T751>Q |
c.2238_2252>GCA(complex) |
12419 |
| 17 |
E19del B |
19 |
p.L747_T751del |
c.2240_2254 del 15 |
12369 |
| 18 |
p.L747_T751del |
c.2239_2253 del 15 |
6254 |
| 19 |
p.E746_T751>V |
c.2237_2252>T |
12386 |
| 20 |
p.E746_S752>I |
c.2235_2255>AAT |
12385 |
| 21 |
p.L747_T751del |
c.2238_2252 del 15 |
23571 |
| 22 |
p.E746_T751del |
c.2236_2253 del 18 |
12728 |
| 23 |
p.E746_S752>A |
c.2237_2254 del 18 |
12367 |
| 24 |
p.E746_S752>V |
c.2237_2255>T (complex) |
12384 |
| 25 |
p.E746_S752>D |
c.2238_2255 del 18 |
6220 |
| 26 |
p.L747_A750>P |
c.2238_2248>GC(complex) |
12422 |
| 27 |
p.L747_E749del |
c.2239_2247 del 9 |
6218 |
| 28 |
p.L747_S752del |
c.2239_2256 del 18 |
6255 |
| 29 |
p.L747_ A750>P |
c.2239_2248 TTAAGAGAAG>C |
12382 |
| 30 |
p.L747_P753>Q |
c.2239_2258>CA(complex) |
12387 |
| 31 |
p.L747_T751>S |
c.2240_2251 del 12 |
6210 |
| 32 |
p.L747_P753>S |
c.2240_2257 del 18 |
12370 |
| 33 |
p.L747_T751>P |
c.2239_2251>C(complex) |
12383 |
| 34 |
p.E746_P753>VS |
c.2237_2257>TCT |
18427 |
| 35 |
S768I |
20 |
S768I |
c.2303G>T |
6241 |
| 36 |
T790M |
20 |
T790M |
c.2369C>T |
6240 |
| 37 |
E20ins A |
20 |
p.D770_N771insG |
c.2310_2311 insGGT |
12378 |
| 38 |
p.P772_H773insTTP |
c.2315_2316 insGACAACCCC |
|
| 39 |
p.P772_H773insGNP |
c.2315_2316 insGGGCAACCC |
|
| 40 |
p.V769_N770insASV |
c.2309_2310 AC>CCAGCGTGGAT |
13558 |
| 41 |
E20ins B |
20 |
p.V769_D770insASV |
c.2307_2308 ins GCCAGCGTG |
12376 |
| 42 |
p.H773_V774insH |
c.2319_2320 insCAC |
12377 |
| 43 |
p.H773L |
c.2318 A>T |
13005 |
| 44 |
p.H773_V774insPH |
c.2319_2320 insCCCCAC |
12380 |
| 45 |
p.V774_C775insHV |
c.2321_2322 insCCACGT |
18432 |
| 46 |
p.D770_N771insSVD |
c.2311_2312 ins GCGTGGACA |
13428 |
| 47 |
L858R |
21 |
L858R |
c.2573T>G |
6224 |
| 48 |
L858R |
c.2573_2574 TG>GT |
12429 |
| 49 |
L861Q |
L861Q |
c.2582T>A |
6213 |
|
(e.g) EGFR Exon21 L858R M & L858R (Genotyping &Sensitivity) |
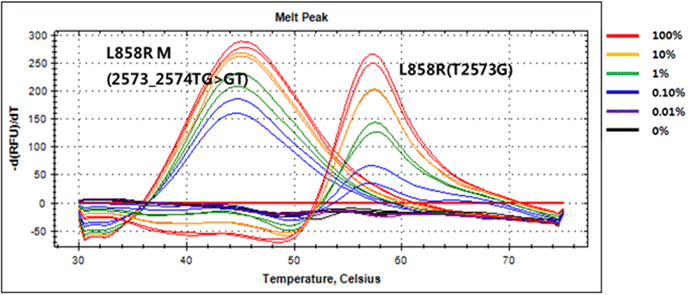 |
Preparation |
EGFR Exon21 L858R M cell line mix |
|
 mutant type: L858R M clone mutant type: L858R M clone
|
|
 wild type: A549 wild type: A549
|
EGFR Exon21 L858R cell line mix |
|
 mutant type:H1975 mutant type:H1975
|
|
 wild type: A549 wild type: A549
|
|
A549 : L858R M clone ratio variation test ⇒ 100%, 10%, 1%, 0.1%, 0.01%, 0 % of L858R M clone
|
|
A549 : H1975 ratio variation test ⇒ 100%, 10%, 1%, 0.1%, 0.01%, 0 % of H1975
|
|
|
(e.g) EGFR Exon20 T790M (Sensitivity) |
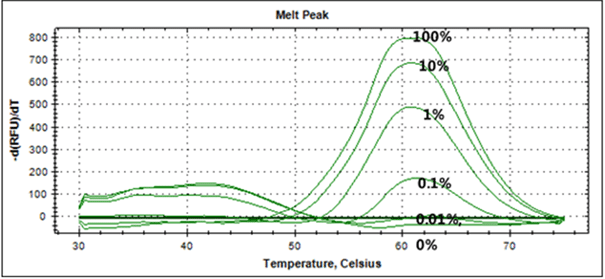 |
Preparation |
EGFR Exon 20 T790M cell line mix |
 mutant type:H1975 mutant type:H1975 |
 wild type: A549 wild type: A549 |
|
A549 : H1975 ratio variation test ⇒ 100%, 10%, 1%, 0.1%, 0.01%, 0 of H1975
|
|
Features |
 Ready-to-use kit with result available within 3 hours. Ready-to-use kit with result available within 3 hours. |
 PNA based mutant enrich genotyping kit- 20 mutations genotyping is possible. PNA based mutant enrich genotyping kit- 20 mutations genotyping is possible. |
 Compatible Real Time PCR - Bio-Rad CFX96 Compatible Real Time PCR - Bio-Rad CFX96 |
 Compatible Sample - Plasma/Serum, FFPE, Fresh tissue Compatible Sample - Plasma/Serum, FFPE, Fresh tissue |
 Test Several Biomarkers in One Plate with same PCR conditions - eg. EGFR+KRAS+NRAS in one plate Test Several Biomarkers in One Plate with same PCR conditions - eg. EGFR+KRAS+NRAS in one plate |
 Excellent Sensitivity - It can be detected less than 0.1% mutant. Excellent Sensitivity - It can be detected less than 0.1% mutant. |
|
|
 |
| 1 |
Hitoshi et al., Peptide nucleic acid-locked nucleic acid polymerase chain reaction clamp-based detection test for gefitinib-refractory T790M epidermal growth factor receptor mutation. Cancer Sci. 99(3): 595-600, 2008 |
|
| 2 |
Minakshi ea al., DISSECT Method Using PNA-LNA Clamp Improves Detection of EGFR T790m Mutation. PLoS ONE. 8(6): e67782, 2013 |
|
| 3 |
Kiyoaki et al., Detection of EGFR Mutations in Archived Cytologic Specimens of Non-Small Cell Lung Cancer Using High-Resolution Melting Analysis. Am J Clin Pathol. 126:608-615, 2006 |
|
| 4 |
Hongdo et al., High resolution melting analysis for rapid and sensitive EGFR and KRAS mutation detection in formalin fixed paraffin embedded biopsies. BMC Cancer. 8(142):1-14, 2008 |
|
|
|